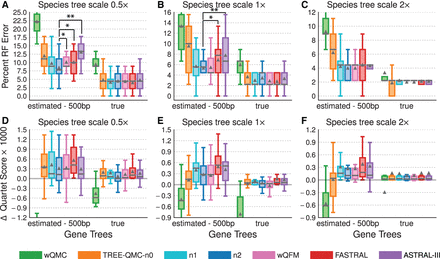
Impact of ILS and GTEE. (A), (B), and (C) Percent species tree error for the avian data set with 1000 estimated or true gene trees and species tree scales 0.5×, 1×, and 2×, respectively. Two-sided Wilcoxon signed-ranked tests were used to evaluate differences between TREE-QMC-n2 versus wQFM, FASTRAL, and ASTRAL3 (nine tests per subfigure). The symbols *, **, and *** indicate significance at P < 0.05, 0.005, and 0.0005, respectively (for 0.5× species tree scale with estimated gene trees, the difference between TREE-QMC-n2 and ASTRAL-II survives Bonferroni multiple comparison correction; see Supplemental Table S6 for details). (D), (E), and (F) show the difference in quartet score between the estimated and true species tree times 1000 for species tree scales 0.5×, 1×, and 2×, respectively (positive values indicate the estimated tree is a better solution to MQSST than the true tree). For species tree heights 0.5×, 1×, and 2×, the ILS level is 60%, 47%, and 35%, respectively, and the GTEE level is 60%, 60%, and 62%, respectively. Results for wQMC are cut off because otherwise the trends cannot be observed (see Supplemental Fig. S2 for full y-axes).











