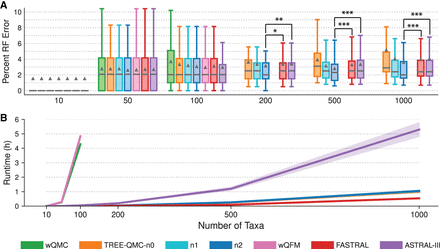
Impact of number of taxa. (A) Percent species tree error across replicates (bars represent medians; triangles represent means; outliers are not shown). The symbols *, **, and *** indicate significance at P < 0.05, 0.005, and 0.0005, respectively (all but one test survives Bonferroni multiple comparison correction; see Supplemental Table S4 for details). (B) Mean runtime across replicates (shaded region indicates standard error). All data sets have species tree height 1×, shallow speciation, and 1000 estimated gene trees. The ILS level is 17%–35%, and GTEE level is 19%–30%. Note: The input processing for wQMC and wQFM does not run within our maximum wall clock time of 18 h for data sets with 200 or more taxa.











