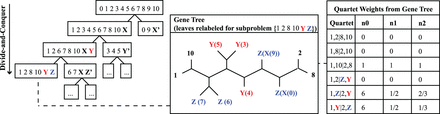
At each step in the divide phase, taxa are split into two disjoint subsets and then artificial taxa are introduced to represent the species on the other side of the split. To compute the quartet weights for a given subproblem, the leaves of each gene tree are relabeled by the artificial taxa. Without normalization (column n0), quartet 1, 2|Y, Z gets zero votes and the alternative quartets get six votes each (note: quartet 1, Y|2, Z gets six votes by taking either species 5, 3, or 4 for label Y and either species 0 or 9 for label Z). With normalization, each gene tree gets one vote for each subset of four labels, although this vote can be split across the three possible quartets. In the uniform normalization scheme (column n1), we simply divide column n0 by the total number of votes cast in the unnormalized case. In the nonuniform normalization scheme (column n2), we leverage that structure implied by the divide phase of the algorithm; the idea is that species should have lesser importance each time they are relabeled by artificial taxa.











