Improving quartet graph construction for scalable and accurate species tree estimation from gene trees
Abstract
Summary methods are widely used to estimate species trees from genome-scale data. However, they can fail to produce accurate
species trees when the input gene trees are highly discordant because of estimation error and biological processes, such as
incomplete lineage sorting. Here, we introduce TREE-QMC, a new summary method that offers accuracy and scalability under these
challenging scenarios. TREE-QMC builds upon weighted Quartet Max Cut, which takes weighted quartets as input and then constructs
a species tree in a divide-and-conquer fashion, at each step forming a graph and seeking its max cut. The wQMC method has
been successfully leveraged in the context of species tree estimation by weighting quartets by their frequencies in the gene
trees; we improve upon this approach in two ways. First, we address accuracy by normalizing the quartet weights to account
for “artificial taxa” introduced during the divide phase so subproblem solutions can be combined during the conquer phase.
Second, we address scalability by introducing an algorithm to construct the graph directly from the gene trees; this gives
TREE-QMC a time complexity of 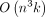 , where n is the number of species and k is the number of gene trees, assuming the subproblem decomposition is perfectly balanced. These contributions enable TREE-QMC
to be highly competitive in terms of species tree accuracy and empirical runtime with the leading quartet-based methods, even
outperforming them on some model conditions explored in our simulation study. We also present the application of these methods
to an avian phylogenomics data set.
, where n is the number of species and k is the number of gene trees, assuming the subproblem decomposition is perfectly balanced. These contributions enable TREE-QMC
to be highly competitive in terms of species tree accuracy and empirical runtime with the leading quartet-based methods, even
outperforming them on some model conditions explored in our simulation study. We also present the application of these methods
to an avian phylogenomics data set.
Footnotes
-
[Supplemental material is available for this article.]
-
Article published online before print. Article, supplemental material, and publication date are at https://www.genome.org/cgi/doi/10.1101/gr.277629.122.
- Received December 23, 2022.
- Accepted May 4, 2023.
This article is distributed exclusively by Cold Spring Harbor Laboratory Press for the first six months after the full-issue publication date (see https://genome.cshlp.org/site/misc/terms.xhtml). After six months, it is available under a Creative Commons License (Attribution-NonCommercial 4.0 International), as described at http://creativecommons.org/licenses/by-nc/4.0/.











