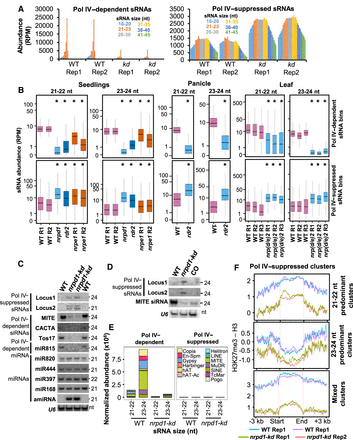
Pol IV complex suppresses sRNA production from several loci. (A) Bar plots showing the abundance of Pol IV–dependent and –suppressed sRNAs of 16- to 45-nt size range. (B) Boxplots showing the abundance of sRNAs from different mutants in rice. sRNAs were size-categorized into 21- to 22-nt and 23- to 24-nt and counted in 100-bp nonoverlapping windows. Plots depict the abundance of sRNAs in each size class over nrp(d/e)2 (leaf), nrpd1 (seedlings), and rdr2 (panicle) dependent and suppressed bins. The data sets were obtained from the NCBI Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/) under accession numbers GSE158709 (seedlings: nrpd1, nrpe1) and GSE130166 (seedlings and panicle: rdr2). The y-axis is scaled to inverse sine hyperbolic function of RPM values. Mann–Whitney U test was used to test statistical significance against WT R1. (*) P-value < 0.0001. (C) sRNA northern blots validating the presence of Pol IV–suppressed sRNAs, –dependent sRNAs, and –independent miRNAs. U6 was used as loading control. (D) sRNA northern blots showing the abundance of sRNAs in WT, kd, and NRPD1 CO lines. U6 was used as loading control. (E) Stacked bar plots showing abundance of Pol IV–dependent and –suppressed sRNAs from transposon categories. (F) Metaplots describing histone H3K27me3 occupancy normalized to total H3 signal over the Pol IV–suppressed sRNA clusters, categorized as 21- to 22-nt predominant clusters, 23- to 24-nt predominant clusters, and the mixed sized clusters.











