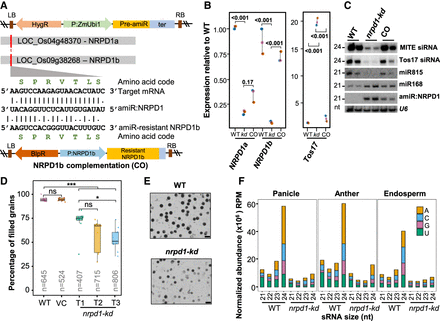
Knockdown of RNA polymerase IV (Pol IV) in rice results in pleiotropic phenotypes. (A) T-DNA map of amiR coding binary construct with the supertransformed NRPD1b complementation (CO) construct. The alignment depicts the amiR binding region and the modifications in the amiR-resistant CDS of NRPD1b that were driven by its cognate promoter (P:NRPD1b). Amino acids encoded by the original and amiR-resistant modifications were unchanged. (BlpR) Bialaphos resistance. The precursor-amiR (Pre-amiR) is driven by maize ubiquitin promoter (P:ZmUbi1). (ter) 35S-poly(A) signal, (HygR) hygromycin selection marker, (RB and LB) right and left border. (B) Plots representing relative abundance of NRPD1a, NRPD1b, and Tos17 transcripts in kd and CO with respect to WT, measured by RT-qPCRs. Student's t-test; P-values mentioned across comparisons. (C) Small RNA (sRNA) northern blots showing the accumulation of 24-nt siRNAs and miRNAs in WT, kd, and CO. U6 was used as loading control. (D) Boxplots showing the percentage of filled grains (n = number of grains). Dots represent the average of panicles in each plant. Tukey's test; (***) P-value of 0.001, (*) P-value of 0.01, (ns) nonsignificant. (VC) Vector control plants that lack amiR encoding region. (E) Representative images of pollen grains of dehisced anther stained with iodine. Scale: 50 μm. (F) Stacked bar plots showing the normalized abundance of sRNAs of different sizes with their 5′ nucleotides. The replicates were merged.











