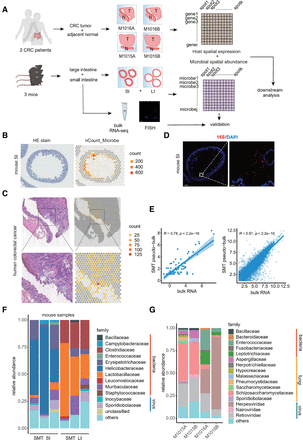
Spatial metatranscriptomics (SMT) of human and murine intestines. (A) A schema of the SMT framework and design of the first part of study. Note that for CRC, tumor and adjacent normal tissue were embedded in the same window and designated T and N, respectively. (B,C) H&E staining of the tissue sections (left) and corresponding spatial feature plot showing microbial sequence per-spot captured (right) for murine small intestine (B) and human colorectal cancer patient tissue, from patient M1016, with zoomed view (C). Representatives are shown, and an additional example can be found in Supplemental Figure S1. (D) FISH result for mouse small intestine section stained for bacteria 16S. Scale bars: 300 μm (left), 20 μm (right). (E) Correlation between SMT-generated pseudobulk measurements (x-axis) and bulk RNA-seq measurements (y-axis) for microbial abundance at genus level (left) and host gene expression (right). Spearman's correlation coefficients and P-values are shown. (F,G) Stacked bar plot showing microbial abundances for mouse (F) and human (G) samples at family level; only the top ones were shown.











