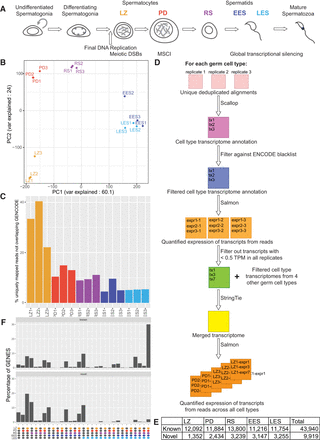
Overview and generation of novel spermatogenic transcriptome. (A) Schematic of spermatogenesis in the adult mouse testis. Key events involving transcriptional regulation are highlighted under the graphic. The following cell types were analyzed in this study: leptotene/zygotene (LZ)- and pachytene/diplotene (P/D)-stage spermatocytes and round (RS), early elongating (EES), and late elongating (LES) spermatids. (B) PCA plot using log2(RPKM) values for genes annotated in GENCODE. (C) Bar plot showing the fraction of uniquely mapped reads that are not overlapping regions previously annotated as genes in GENCODE. (D) Schematic overview of the de novo transcriptome assembly approach. (E) Table showing number of genes found in each cell type. Known indicates any overlap with a previously GENCODE annotated gene, and novel genes do not possess any such overlap. (F) Bar plot showing expression patterns of known and novel genes in spermatogenesis. Colored balls under plot indicate cell type(s) in which the genes are expressed (with gray representing not expressed).











