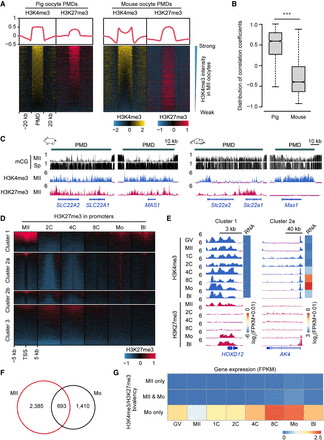
H3K4me3/H3K27me3 bivalency in porcine oocytes and early embryos. (A) Heat maps and aggregate profile plots showing the H3K4me3 and H3K27me3 enrichment in porcine and mouse MII oocytes at their oocyte PMDs. PMDs were ordered descending by their H3K4me3 intensities in MII oocytes. (B) Box plot showing the distribution of Pearson's coefficients between H3K4me3 and H3K27me3 intensities in porcine and mouse MII oocytes at their oocyte PMDs. (***) P < 0.001 by Wilcoxon test. (C) Genome browser snapshots showing the orthologous genomic regions co-occupied H3K4me3 and H3K27me3 in porcine oocyte PMDs, but did not co-occupy these modifications in mouse oocyte PMDs. DNA methylation levels (mCG) in oocyte and spermatozoa are also shown. (D) Heat maps showing the H3K27me3 dynamics at all gene promoters (TSS ± 5 kb) in porcine oocytes and early embryos. Gene clusters were defined in Figure 2A. Gene promoters in each cluster were ordered descending by their H3K27me3 intensities in MII oocytes. (E) Genome browser snapshots showing the H3K4me3 and H3K27me3 dynamics and transcriptional levels (FPKM) of candidate genes for Cluster 1 and Cluster 2a in porcine oocytes and early embryos. (F) Venn diagram showing the oocyte-specific (MII), morula-specific (Mo), and overlapping bivalent gene numbers between porcine MII oocytes and morulae. (G) Heat map showing the transcriptional levels (FPKM) of oocyte-specific (MII only), morula-specific (Mo only), and overlapping (MII & Mo) bivalent genes in porcine oocytes and early embryos.











