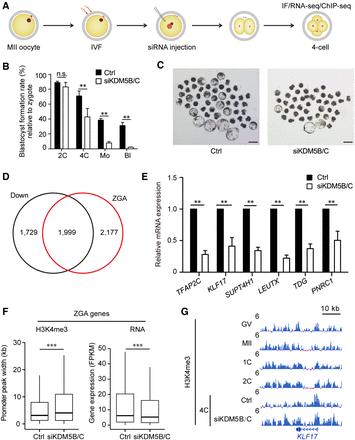
Knockdown of KDM5B and KDM5C disrupts porcine ZGA and embryonic development. (A) Experimental design of KDM5B and KDM5C knockdown in porcine IVF embryos. (B) Bar plot showing the blastocyst formation rates in KDM5B and KDM5C knockdown (siKDM5B/C) and control (Ctrl) groups when compared to the harvested zygotes. Error bars represent the SD. (n = 3, 100 oocytes per repeat.) (**) P < 0.01 by two-tailed t-test, (n.s.) no significance. (C) Microscopy images showing the blastocyst formation in siKDM5B/C and control groups. Scale bar, 200 μm. (D) Venn diagram showing that over half (1999/3728) of down-regulated genes in siKDM5B/C four-cell embryos are overlapped with porcine ZGA genes. (E) Bar plot showing the relative mRNA expression of six porcine ZGA genes in siKDM5B/C and control embryos at the four-cell stage. Error bars represent the SD. (n = 3, 30 embryos per repeat). (**) P < 0.01 by two-tailed t-test. (F) Box plots showing the width of H3K4me3 peaks at ZGA gene promoters and the transcriptional levels (FPKM) of ZGA genes in siKDM5B/C and control groups. (***) P < 0.001 by Wilcoxon test. (G) A genome browser snapshot showing the H3K4me3 enrichment at KLF17 location in siKDM5B/C and control embryos at the four-cell stage.











