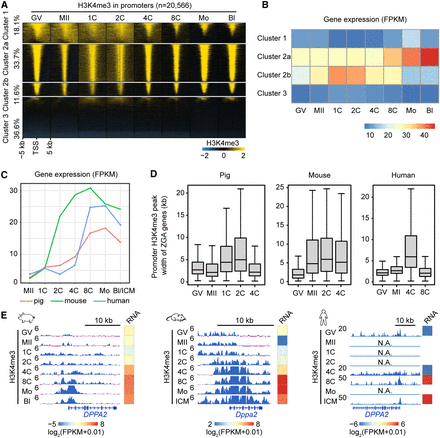
De novo H3K4me3 establishment at ZGA gene promoters upon fertilization. (A) Heat maps showing the H3K4me3 dynamics at all gene promoters (transcriptional start site [TSS] ± 5 kb) in porcine oocytes and early embryos. k-means analysis (k = 3) was performed to classify these promoters into three clusters based on their H3K4me3 intensities in GV oocytes, in which Cluster 2 could be further divided into two subclusters based on their H3K4me3 intensities in zygotes. Gene promoters in each cluster were ordered descending by their H3K4me3 intensities in GV oocytes. (B) Heat map showing the transcriptional levels (average fragments per kb per million reads [FPKM]) of genes in each cluster in porcine oocytes and early embryos. (C) Line plots showing the transcriptional levels (FPKM) of porcine, mouse, and human zygotic genome activation (ZGA) genes in their oocytes and early embryos. (D) Box plots showing the peak width of promoter H3K4me3 for porcine, mouse, and human ZGA genes in oocytes and early embryos. (E) Genome browser snapshots showing the H3K4me3 dynamics and transcriptional levels (FPKM) of orthologous ZGA genes, Dppa2/DPPA2, in porcine, mouse, and human oocytes and early embryos. Broad H3K4me3 was established at ZGA gene promoters in porcine and human pre-ZGA embryos (pig: 1C-2C; human: 4C) and reshaped into sharp peaks in their peri-ZGA embryos (pig: 4C-8C; human: 8C).











