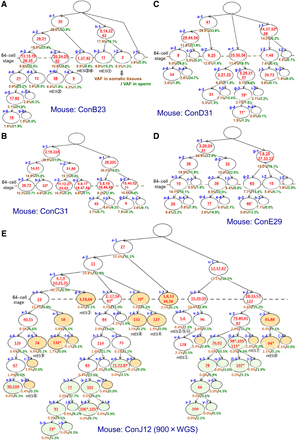
Reconstructed lineage trees based on de novo mutations occurring during the first several postzygotic divisions. Reconstructed lineage trees of five animals based on the mosaic mutations detected through whole-genome sequencing (WGS) with 100× (A–D) and 900× (E) coverages. The red numbers represent newly arising mutations. The underlined numbers represent mutations also observed in the offspring. Numbers with an asterisk represent mutations added using the genotyping results of the offspring. The numbers with a hash mark represent mutations with a GERP score of more than three (expected to have a harmful effect). The nuclear transfer embryonic stem (ntES) below the cell positions represents the lineages leading to each ntES cell line established in the mice ConB23 (A) and ConJ12 (E). The brown and green numbers (%) below each cell position represent the VAFs of labeled cells’ autosomal mutations (or the estimated values if there are no mutations) in somatic tissue averages and in a sperm sample, respectively. In the ConJ12 tree (E), in sperm or all somatic tissue samples, cells with VAF values comparable to the background were colored orange and light green as somatic cell–specific lineages and germ cell–specific lineages, respectively.











