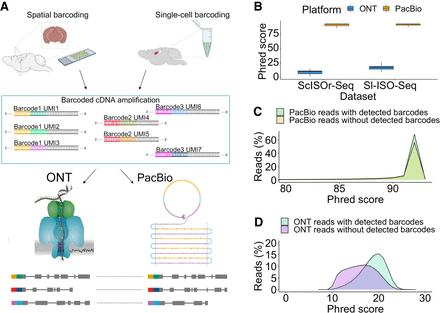
Outline and primary read characteristics. (A) Individual reverse transcription events turn an individual RNA molecule into a barcoded RNA–cDNA hybrid, which is amplified into many cDNA molecules that carry the same barcode and UMI. We previously performed this process in two distinct ways: by single-cell 10x Genomics barcoding and by spatial 10x Genomics Visium barcoding. Aliquots of these cDNAs are then sequenced on PacBio and ONT. Using the identity of barcode and UMI, we can detect individual RNA molecules whose cDNA copies have been sequenced on both ONT and PacBio. We refer to these read pairs as RT read pairs. (B) Comparison between Phred scores of PacBio CCS and ONT reads from both data sets. (C) Phred score distribution for PacBio CCS reads from the Sl-ISO-Seq data set with (light green) and without (yellow) detected barcodes. (D) Phred score distribution for ONT reads from the Sl-ISO-Seq data set with (light blue) and without (purple) detected barcodes.











