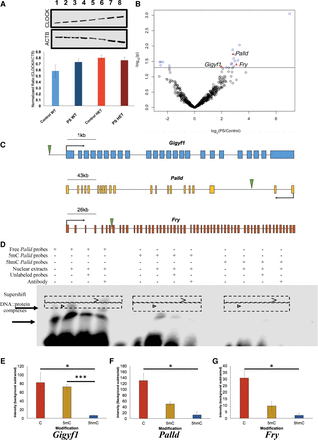
Functional assays reveal that 5hmC represses CLOCK binding. (A) Western blot analysis identified no gross differences in CLOCK protein expression between Control-WT (lanes 1,2), PS-WT (lanes 3,4), Control-HET (lanes 5,6), or PS-HET (lanes 7,8; top panel) or in ACTB (middle panel), used as a loading control in all lanes. A bar plot (bottom panel) depicts the quantified levels of CLOCK proteins normalized to ACTB from each respective lane. Error bars show the SEM (N = 2). (B) A volcano plot displays the –log2(fold-change) of PS-HET compared with Control-HET (x-axis) and the significance (y-axis) from differential analysis of ChIP-seq data. Open circles (blue/black) and red triangles represent DhMRs containing a canonical CLOCK binding site. DhMRs with significant differential binding of CLOCK (blue open circles and red triangles) are located above the significance line (P-value < 0.05). Red triangle DhMRs also show differential gene expression (Gigyf1, Palld, and Fry). (C) Gene schematics of Gigyf1 (blue), Palld (yellow), and Fry (orange) are shown and indicate the location of the DhMRs containing the differential CLOCK binding (green triangle). (D) Electromobility shift assays with hippocampal lysates. Lysate shift (lanes 2,6,10), cold competition (lanes 3,7,11), and supershift (lanes 4,8,12) of DNA probes corresponding to a CLOCK E-box motif in the DhMR of Palld that showed differential CLOCK binding. Biotinylated DNA probes contained either an unmodified E-box motif (lanes 1–4), a methylated E-box motif (lanes 5–8), or a hydroxymethylated E-box motif (lanes 9–12). Regions containing bands of interest are highlighted (dashed boxes); DNA::protein complexes that are abrogated with cold competition are indicated (black arrowheads) and supershift complexes are shown (carrot). (E–G) Quantifications of biotinylated DNA probe intensities. The E-box modifications (x-axis) and mean biotinylated DNA probe intensity (y-axis) are depicted ±SEM for Gigyf1 (E), Palld (F), and Fry (G) probes. The asterisk (*) denotes a significant decrease in CLOCK binding box: (*) P-value < 0.05; (***) P-value < 0.001.











