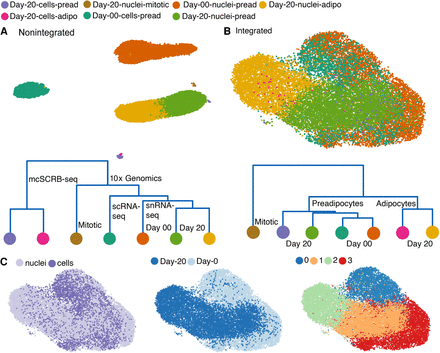
Integration of the snRNA-seq and scRNA-seq data sets. (A, top panel) UMAP visualization of nonintegrated scRNA-seq and snRNA-seq data sets for both white preadipocyte (day 0) and mature adipocyte (day 20), for a total of four batches (total 18,717 barcodes). (Bottom panel) Cluster dendrogram for nonintegrated data sets based on the eigenvalue-weighted Euclidean distance matrix constructed in latent-dimension space inferred using scVI. (B) UMAP visualization and cluster dendrogram of scRNA-seq and snRNA-seq data sets as in A after integration using scVI tools (total 18,717 barcodes). See also Supplemental Note S4 and Supplemental Figure S10. (C) UMAP visualization of scVI integrated data set with barcodes annotated by sequencing technique (left), harvest day (middle), and joint unsupervised clustering (right).











