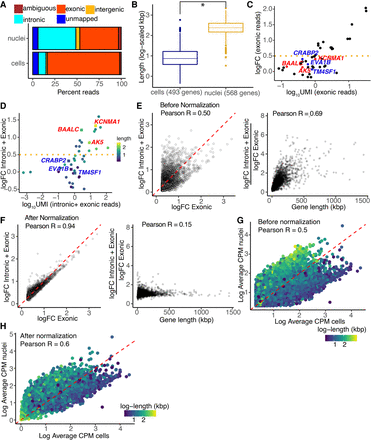
Gene length–associated detection bias in the nuclear transcriptome. (A) Distribution of reads in the scRNA-seq and snRNA-seq preadipocyte data sets. (B) Distribution of gene length for genes enriched in cells (in blue) and nuclei (in yellow) with log fold change > 1 and FDR < 0.05, including both intronic and exonic reads. (C) logFC versus log-UMI counts in white nuclei when using only exonic reads, where each dot represents a white preadipocyte–enriched gene (white vs. brown DE test) detected using the scRNA-seq data set. Horizontal dotted line indicates a logFC cutoff value of 0.5 used as a threshold for DE testing. All genes had a logFC > 0.5 in the scRNA-seq data set. (D) logFC versus log-UMI counts in white nuclei when using both intronic and exonic reads. Each dot is the same as in panel C. Horizontal dotted line indicates a logFC cutoff value of 0.5 used as a threshold for DE testing. Highlighted genes are represented with a square symbol. (E, left): logFC for nuclear-enriched genes when using only exonic reads or both intronic and exonic reads before normalization. Each dot represents a gene enriched in nuclei using exonic-only reads with logFC > 0.25 and FDR< 0.05. Red dotted line indicates y = x axis. (Right) Ratio of y-axis value over x-axis value for genes in left panel, plotted as a function of their length. (F, left) logFC for nuclear-enriched genes when using only exonic reads or both intronic and exonic reads after normalization. Each dot is the same as in panel E. Red dotted line indicates y = x axis. (Right) Ratio of y-axis value over x-axis value for genes in left panel, plotted as a function of their length. (G,H) Average expression of genes in white cells and white nuclei when using both intronic and exonic reads, without normalization (G) and with normalization (H). For normalization strategy, see Supplemental Note S5. Each dot represents a gene with average counts per million (CPM) >1 when using both intronic and exonic reads, without normalization, in both cells and nuclei. White nuclei were randomly selected to have as many barcodes as white cells. Red dotted line has a slope of 1.











