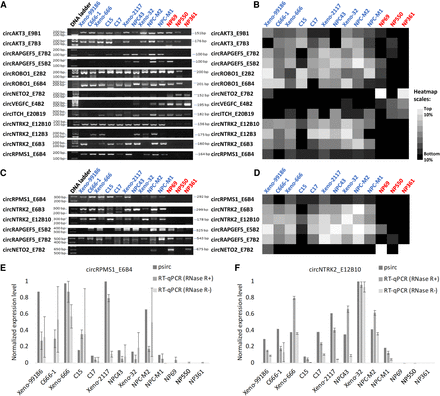
Experimental validations of the computational results of psirc. (A) Validation of differentially expressed BSJs in the RNase R–treated NP and NPC RNA samples using RT-PCR. Each row corresponds to a BSJ and each column corresponds to a sample, with the NPC samples labeled in blue and the NP samples labeled in red. (B) Expression levels of the same BSJs determined by psirc. The expression value of each BSJ was computed by summing up the TPM values of all transcripts that involved this BSJ. (C) Validation of differentially expressed full-length transcript isoforms using RT-PCR. (D) TPM values of the same full-length isoforms inferred by psirc. (E,F) RT-qPCR results of two short full-length transcript isoforms, circRPMS1_E6B4 (E) and circNTRK2_E12B10 (F), with or without RNase R treatment. Each RT-qPCR experiment was repeated three times with independent RT preparations, with the standard deviation of each triplicate indicated by the error bars. In each of these two panels, normalization was performed by dividing each value by the largest value among all samples.











