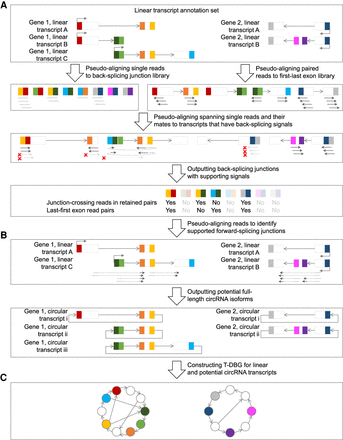
The psirc method. In the first step (A), sequencing reads are pseudoaligned to potential BSJs in single-end mode and to the first and last exons of each transcript in paired-end mode. For each read that is pseudoaligned to a BSJ (by means of rotating the end of the downstream exon to the beginning of the upstream exon), if its mate read is not pseudoaligned to the same transcript or not pseudoaligned in the opposite orientation, both of them will not be considered to support the BSJs (arrows in light gray with a red cross next to them). In the second step (B), the potential circRNA transcript isoforms are determined as those with all forward-splicing and backward-splicing junctions supported by sequencing reads. In the third step (C), a T-DBG is constructed for all linear and circular transcript isoforms for estimating their expression levels by another round of pseudoalignment.











