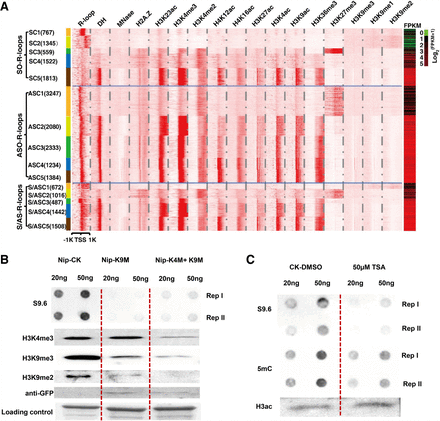
Epigenomic signatures associated with R-loop overlapping genes. (A) k-Means clustering analysis of read densities (normalized read counts) calculated with a 50-bp sliding window using sequencing data from DRIP-seq, ChIP-seq, DNase-seq, or MNase-seq approaches (where k = 5). The resulting profiles of epigenomic features of non-TE genes are displayed as a heatmap; region shown is ±1 kb of the TSSs. Non-TE genes were organized based on the types of R-loops present: SO-R-loops, ASO-R-loops, and S/AS-R-loops. The number of genes in each cluster is listed in brackets immediately after the cluster name; expression levels of genes (FPKM values) involved are shown in the rightmost lane. (B) Dot and western blotting assays using antibodies as shown at the left side. Each sample had two technical repeats per blot for the dot blotting but only one replicate for the western blotting. The experiment was repeated twice with similar results being obtained. The three wells from left to right in the row of “Loading control” are Coomassie Blue stained and contain the same amount of total proteins extracted from the Nip, Nip-K9M, and Nip-K4M + K9M lines. (C) Dot and western blotting assays used antibodies as shown. Each DNA sample had two technical repeats per blot, and each experiment was repeated two times and the results were stable. Dot signals or western bands were cut from original blots that can be found in Supplemental Fig. S10.











