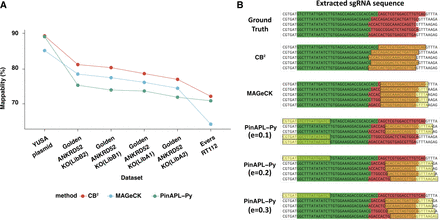
CB2 outperforms MAGeCK and PinAPL-Py in the percentage of mapped reads over six benchmark data sets. (A) Read mappability of CB2, MAGeCK, and PinAPL-Py across six different data sets. (B) Representative examples of reads that were not mapped by MAGeCK or PinAPL-Py. Adapters are highlighted with green; sgRNAs, with red. Yellow boxes show the predicted locations of sgRNAs by each method. Several parameters were used to optimize performances of PinAPL-Py.











