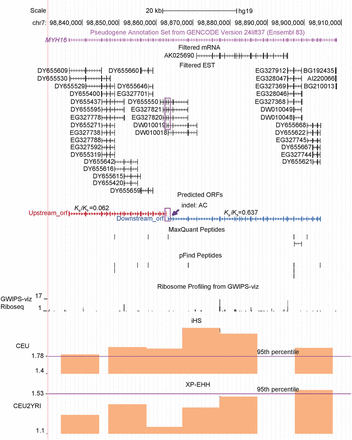
Gene structure and expression of MYH16. The UCSC Genome Browser snapshot around the MYH16 locus is presented. The tracks including “Filtered mRNA” and “Filtered EST” show only the entries that are uniquely mapped to this locus. Because these mRNA and ESTs share a single compatible exon–intron structure, one transcript encoding 43 exons can be inferred, as shown by the GENCODE (Ensembl) pseudogene annotation track. MYH16 can therefore be translated as two ORFs separated by the indel (a deletion of “AC”), whose position is highlighted by the purple arrow. Two continuous exons supported by four ESTs (e.g., DY655550) are highlighted by a purple frame, indicating that these two ORFs are encoded by a single transcript. Uniquely mapping peptides detected by two algorithms (MaxQuant, pFind) and Ka/Ks values reported by Perry et al. (2005) are shown. The iHS and XP-EHH tracks from PopHuman were added, with the 95th percentile indicated by the purple line. CEU and YRI represent one European population and one African population, respectively.











