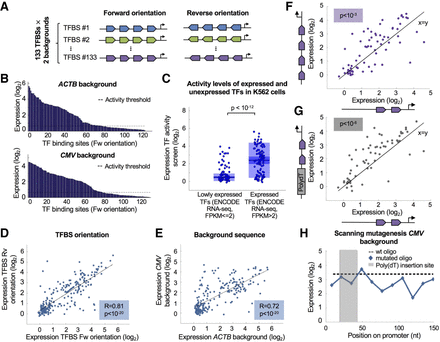
TF activity screen for 133 binding sites and the effect of nucleosome disfavoring sequence on expression. (A) Illustration of designed oligos for TF activity screen. One hundred thirty-three binding sites for 70 TFs were placed in four copies in either the forward or the reverse orientation in two backgrounds. (B) Expression measurements of oligos containing forward TF binding sites in two different backgrounds. Each bar represents a single binding site. Activity threshold determined by the empty vector is denoted. (C) TF activity measurements of expressed and unexpressed TFs as determined by ENCODE RNA-seq in K562 cells. Low activity is obtained for unexpressed TFs (P < 10−12, Wilcoxon rank-sum test). (D) Comparison between expression measurements of binding sites in two orientations. Each dot represents a pair of sequences for the same binding site placed in the forward or the reverse orientation (R = 0.81, P < 10−20, Pearson's correlation). (E) Comparison between expression measurements of binding sites in different backgrounds. Each dot represents a pair of sequences for the same binding site placed in the ACTB or the CMV backgrounds (R = 0.72, P < 10−20, Pearson's correlation). (F) Testing the effect on expression of adding two TF binding sites. Each dot represents a pair of designed promoters with either two or four sites for one of the 70 TFs tested in the CMV background. An increase in expression is observed for most TFs (P < 10−3, Wilcoxon signed-rank test). (G) Testing the effect on expression of nucleosome disfavoring sequence. A 25-mer poly(dA:dT) tract was added upstream to two binding sites for 70 TFs. An increase in expression is observed for most TFs (P < 10−6, Wilcoxon signed-rank test). (H) Systematic scanning mutagenesis to identify cis-regulatory elements in the CMV promoter. Eleven mutated oligos were designed; each contains a 14-nt window in which all nucleotides were mutated. Each dot represents expression of one mutated oligo. No elevation in expression is observed when mutating the sequences in which the poly(dA:dT) was inserted.











