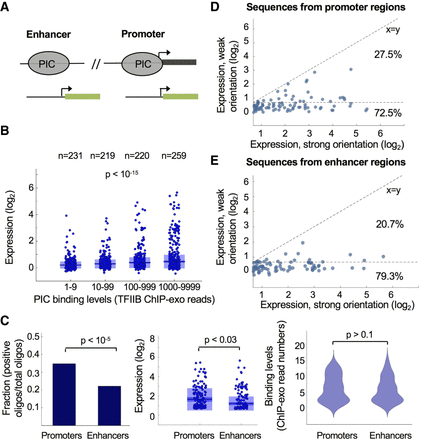
Functional measurements of autonomous core promoter activity of PIC binding sequences from promoters and enhancers. (A) Illustration of the designed sequences matching 508 PIC binding regions in promoters and enhancers that were identified by ChIP-exo measurements in K562 cells (Venters and Pugh 2013). (B) Comparison between core promoter activity of sequences with different PIC binding levels (TFIIB ChIP-exo). Data were binned into four groups according to the number of ChIP-exo reads, and expression measurements were compared between bins (P < 10−15, Kruskal-Wallis test). (C) Comparison between the fraction of positive core promoters for PIC binding sequences from promoters and enhancers (left; P < 10−5, two-proportion z-test) and the activity levels of positive sequences from both groups (middle; P < 0.03, Wilcoxon rank-sum test). To avoid biases in activity stemming from different PIC binding levels, sequences with the same number of ChIP-exo reads were selected in the design process (right; P > 0.1, Wilcoxon rank-sum test). (D,E) Comparison between core promoter activity of PIC binding sequences from promoters (D) and enhancers (E) in two orientations. Each dot represents a distinct PIC binding site that presented positive activity in at least one orientation. Expression measurements of designed sequences are shown for the stronger and weaker orientations of each pair of sequences. The horizontal dashed line represents the activity threshold as determined by empty vector measurements; the diagonal dashed line, a theoretical x = y line expected for promoters with equal expression in the two orientations.











