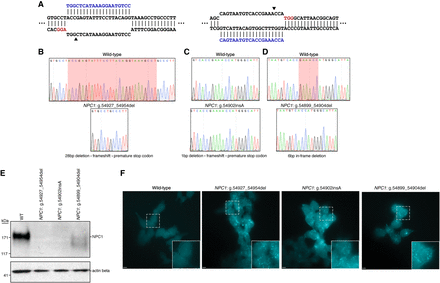
Generation and characterization of three haploid models of Niemann–Pick disease type C (NPC). (A) Diagrams illustrating the two targeted sites in NPC1. Arrowheads indicate the predicted DSB site. (B–D) Sequencing chromatographs showing wild-type NPC1 (top) and the specific disruption in each isogenic edited cell clone (bottom). (B,D) Red highlighted region indicates the locations of the deletions in edited clones. (C) Red highlighted region indicates the position of the insertion in the edited clone. (E) Western blot analysis from total protein lysate from wild-type HAP1 cells and the three edited cell clones illustrating absent or reduced NPC1 protein expression. Actin beta was used as a loading control. (F) Filipin staining reveals deposits of intracellular cholesterol in edited cells that are absent in wild-type cells. White dashed-bordered box has been enlarged twofold and inset at bottom right. Scale bars, 6.3 μm.











