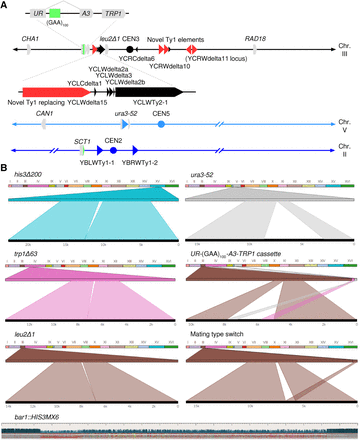
Known deletions and rearrangements in SMY502 versus S288C. (A) Map of Chromosomes II, III, and V, indicating positions of genes, centromeres, and Ty elements. Diagonal lines represent contiguous sequences not in display, such that the displayed portions are pictured to scale. Genes are shown by gray boxes with points indicating the orientation. Centromeres are marked by circles. Ty elements are indicated by triangles, with black indicating Ty elements found in the S288C reference genome, and red indicating previously unannotated Ty elements. The point of the triangle indicates orientation. Small black triangles represent solo LTR sequences, also known as delta elements. Zoomed-in portions of Chromosome III show a cluster of Ty elements, as well as the UR-(GAA)100-A3 selectable cassette. Bright green portions represent the location of (GAA)n repeats. (B) Ribbon single-read views highlighting known large-scale genomic changes in the SMY502 parent strain, mapped to the S288C reference genome. For each panel, the top bar contains a color-coded list of chromosomes, whereas the bottom black bar displays the full sequencing read. Windows connect the portion of the read that maps to the chromosomal position. his3Δ200, trp1Δ63, and leu2Δ1 are 1- to 2-kb deletions and are observed as split reads in which part of the reference sequence (top) is missing from the read (bottom). Ura3-52 is an insertion of an ∼6-kb Ty element, observed as a split read in which sequence not in the reference (top) is found in the read (bottom). Because of high sequence similarity among Ty elements, the insertion is not always associated with a particular part of the reference sequence in each individual read. Our UR-(GAA)100-A3-TRP1 selectable cassette also maps as an insertion, but the 5′ portion of URA3 and the TRP1 gene are both matched to their respective genomic locations in the reference sequence. The difference in mating types between S288C and our strain is also observed as a split read, due to the peculiar control of yeast mating type, in which one of two inactive regions on either end of Chromosome III is copied via HR into the active mating loci, located near the middle of Chromosome III. One allele, bar1::HIS3MX6, was not apparent in the Ribbon analysis, because the BAR1 gene was replaced with a similarly sized marker gene. The HIS3MX6 marker was aligned to the reference with a number of mismatches and short gaps, which was readily apparent in the UGENE alignment. In this view, gray boxes indicate bases that match the reference sequence, and colored boxes indicate bases that do not match the reference sequence: (blue) G; (green) C; (yellow) A; (light red) T; (dark red) deletion. The blue bars above represent read depth at each position.











