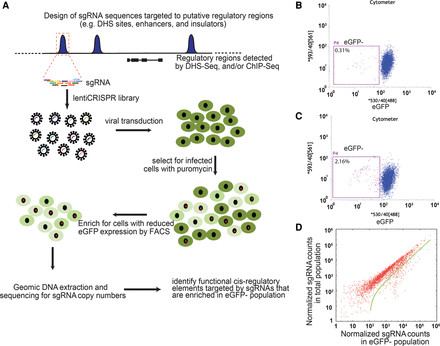
Experimental design of a high-throughput CRISPR/Cas9-mediated screen for identifying cis-regulatory elements. (A) Workflow of lentiCRISPR screening strategy to identify functional regulatory elements. sgRNA library sequences were designed to create random mutations at 174 predicted regulatory regions (blue peaks) in the POU5F1 TAD via nonhomologous end joining (NHEJ). sgRNA sequences were synthesized in an array-based oligo pool, cloned into the lentiCRISPR plasmid, and packaged into lentiviral libraries. We then used lentiviral libraries to infect H1 POU5F1-eGFP cells to generate random mutagenesis of the 174 candidate regions. hESCs with mutations at cis-regulatory elements affecting POU5F1 expression can be identified as eGFP− cells. (B) Control H1 POU5F1-eGFP without lentiviral infection were dissociated into single cells and subjected to FACS analysis to determine the eGFP− (P4) gate for eGFP− population; 0.31% indicates the ratio of P4 in parental live singlets. (C,D) The H1 POU5F1-eGFP cells were infected with lentiCRISPR library by spin infection at low multiplicity of infection (MOI). Twenty-four hours after infection, the cells were cultured for 7 d under puromycin selection; for another 10 d, without puromycin. (C) The cells were subjected to FACS analyses. (2.16%) The ratio of P4 in parental population. (D) The eGFP− P4 population was collected by FACS sorting. Genomic DNA was isolated from P4 and nonsorted control cells, followed by PCR amplification of sgRNA sequence and deep sequencing. Scatter plot for sgRNA read counts in eGFP− cells compared with the control cells after LOESS normalization. Dots underneath the green line are sgRNAs with at least twofold enrichment in the eGFP− cells compared with the control population.











