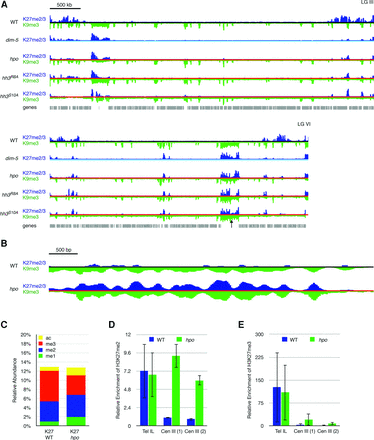
Loss of HP1 leads to H3K27me2/3 redistribution. (A) ChIP-seq tracks of H3K27me2/3 (dark blue) and H3K9me3 (inverted; green) are displayed for LG III and LG VI in a wild-type strain (black line), a dim-5 strain (light blue line), an hpo strain (red line), and two histone mutants, hh3R8A and hh3S10A (both indicated by red lines). H3K9me3 is relatively unaffected by a deletion of hpo, while H3K27me2/3 is redistributed to constitutive heterochromatin in an hpo strain, as in the dim-5 strain. Each of the histone mutants, which largely lose HP1 localization and have reduced H3K9me3, shows H3K27me2/3 redistribution similar to the dim-5 strain. The scale bar (500 kb) at the top left applies to both chromosomes. The black arrow indicates a centromere region on LG VI that is shown expanded 1000-fold below. (B) The distributions of H3K27me2/3 (blue) and H3K9me3 (inverted; green) were examined at a window size of 25 bp in the wild-type (black line) and hpo (red line) strains. At this scale (bar, 500 bp), the peaks of H3K9me3 and H3K27me2/3 in the hpo strain closely resemble one another, which is consistent with both histone marks occupying the same nucleosome. (C) The relative abundance of four modifications (mono- [me1], di- [me2], and trimethylation [me3] and acetylation [ac]) on the H3K27 residue as determined by LC-MS/MS is shown for wild-type and hpo strains. (D) ChIP-qPCR to detect the relative enrichment of H3K27me2 in wild-type and hpo strains at subtelomere IL (Tel IL) and two centromere III loci (Cen III). (E) The relative enrichment of H3K27me3 in wild-type and hpo strains at the same three loci as in D.











