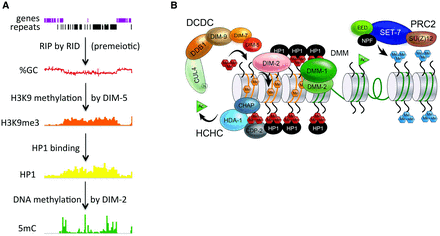
Heterochromatin formation in Neurospora crassa. (A) An ∼500-kb region, including the centromere of LG III, is shown to illustrate the pathway leading to constitutive heterochromatin in Neurospora. During the sexual phase of the life cycle, the genome defense system RIP (repeat-induced point mutation) recognizes repeated DNA (e.g., transposons and other repeated DNA shown as black rectangles) and litters them with C:G-to-T:A mutations (Selker 1990). In vegetative cells, lysine 9 of histone H3 (H3K9) associated with G:C-poor DNA is methylated by DIM-5, generating H3K9me3 (orange track), which is bound by the HP1-DIM-2 complex (yellow track) and catalyzes DNA methylation (green track). Perturbation of any step in the pathway eliminates the downstream steps without significantly influencing earlier steps. (B) Key proteins required to form constitutive (left) or facultative (right) heterochromatin. Methylation of H3K9 by DIM-5 depends on all five members of the DCDC (DIM-5/-7/-9, CUL4/DDB1 complex) (Lewis et al. 2010a). HP1 directly binds to H3K9me3 (cluster of three red hexagons labeled “Me”) and recruits the DNA methyltransferase DIM-2 (Honda and Selker 2008), resulting in a genome-wide correlation between H3K9me3 and DNA methylation (orange hexagons labeled “Me”) (Lewis et al. 2009). HP1 is also involved in the HCHC deacetylase silencing complex (Honda et al. 2012) and the DMM (Honda et al. 2010) complex, which limits spreading of constitutive heterochromatin. Methylation of H3K27 (cluster of three blue hexagons labeled “Me”) is carried out by the PRC2 complex consisting of SET-7, EED, SU(Z)12, and NPF (Jamieson et al. 2013).











