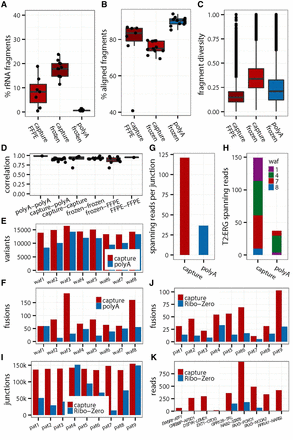
Assessment of capture transcriptomes from clinical frozen and FFPE samples. (A–C) Comparative analysis of paired capture and poly(A) libraries (grouped by patient) derived from FFPE blocks and frozen tissue: (A) efficiency of rRNA depletion; (B) alignment rates; (C) fragment diversity (FD)—a compound measure of transcriptome quality sensitive to coverage, complexity, and insert size; more complex and well-covered libraries have higher FD values. (D) Within patient correlation of gene expression (log2[cpm]) by library type (poly(A) vs. capture) and source material (frozen vs. FFPE). (E,F) Sensitivity of libraries for detecting genetic changes by patient from frozen libraries: (E) number of called variants; (F) number of called candidate fusions. (G,H) Robustness of fusion detection: (G) average read support per fusion; (H) number of supporting reads for each cohort patient with the TMPRSS2-ERG fusion detected. (I,J) Paired capture and Ribo-Zero libraries from FFPE: (I) number of detected splice junctions; (J) number of called candidate fusions. (K) Selected candidate oncogenic fusion for each patient (read support).











