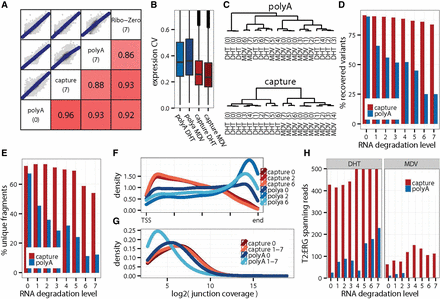
Improved performance of exome-capture transcriptomes from low quality RNA samples. (A) Correlation of absolute levels of gene expression (log2[cpm]) between a reference library from intact RNA (poly[A] level 0) and libraries from degraded RNA (level 7). (B) Impact of RNA degradation on gene expression accuracy measured as the average coefficient of variation (CV)—larger values indicate more variable measurements. (C) Impact of expression accuracy on the unsupervised clustering of samples with biological differences confounded by technical variation. (D) Sensitivity of detection of single nucleotide variants in libraries of varying RNA quality. (E) Library complexity estimated as the percentage of unique (nonduplicate) fragments among all counted fragments. (F,G) Assessments of uniformity of transcript coverage. (F) Smooth density estimate of read start positions along the scaled gene bodies (genes <10 kb were excluded). (G) Distribution of splice junctions by depth of coverage. (H) Sensitivity of detecting the TMPRSS2-ERG fusion (junction coverage).











