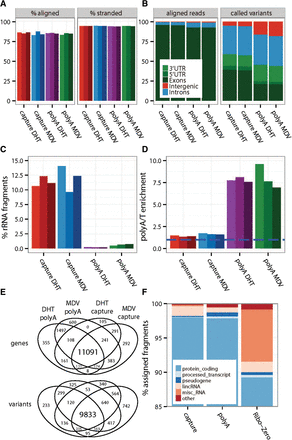
Similarity of poly(A) and capture transcriptomes from intact RNA. Properties of fragments from both types of libraries. Separate bars (colors) for each replicate in A,C,D. (A) Alignment rates and library strand-specificity (% fragments aligned to the transcribed strand). (B) Types of genomic alignment regions by fraction of assigned fragments and fraction of discovered variants. (C) Efficiency of rRNA depletion (% fragments aligning to ribosomal RNA). (D) Overrepresentation of poly(A) and poly(T) hexamers. (E) Global concordance of detected genes and called variants within all exonic regions. (F) Fraction of assigned reads by biological gene category.











