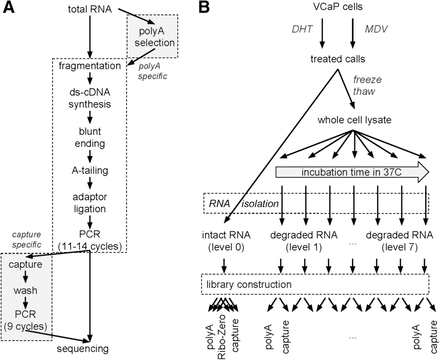
The exome-capture transcriptome protocol. (A) Flow-chart of library preparation protocols. Steps unique to each protocol are highlighted. Enrichment for mRNA occurs at the RNA or cDNA stage, respectively, for poly(A) and capture RNA-seq. (B) Controlled in vitro degradation through cell lysis and warm incubation. VCaP cells were treated with DHT or MDV3100. Intact RNA, RNA integrity number (RIN) 10, was extracted, and libraries were prepared in technical triplicates. In parallel, RNA was degraded by warm incubation for increasing amounts of time. Paired poly(A) and capture libraries were prepared from the same RNA at each degradation level.











