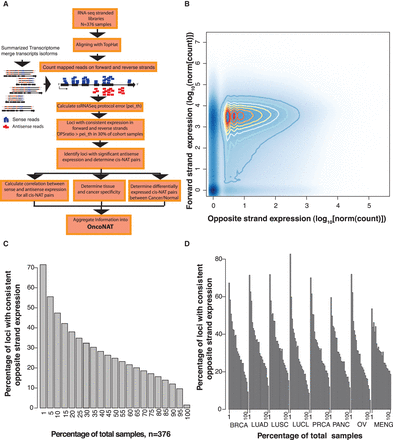
Antisense expression is pervasive across the human transcriptome. (A) Bioinformatics workflow for characterization of expressed antisense loci. The summarized transcriptome was built by reconstructing the longest high quality annotation for each gene, using transcript and exon information provided in the Ensembl v69 assembly. This procedure generated 42,124 gene models (Supplemental Data S4) whose expressions on forward and opposite strands were calculated (Supplemental Data S5, S6). The protocol error rate of a library (pei) corresponds to the fraction of reads mistakenly mapped to the opposite strand but generated by the forward gene (Methods). Bona fide antisense transcripts are loci that have an OPSratio > pei_th in at least 30% of the cohort samples and have a statistically significant antisense expression according to NASTI-seq (Methods). This bioinformatics workflow nominates expressed antisense loci across cancer subtypes, establishes their pattern of expression, and generates a catalog of tumor suppressor and oncogenes with significant antisense expression, OncoNAT (Supplemental Data S11–S15). (B) Density plot of forward and opposite strand expression. Expression over forward and opposite strands is calculated for each locus per sample. Expression in the opposite strand is in general two to three orders of magnitude lower (median of opposite/forward = 0.001) than the expression of the forward strand. (C) The percentage of loci with consistent expression in the opposite strand. A loci is considered to have consistent opposite strand expression if the OPSratio > pei_th in at least 30% of the samples in the cohort. (D) The percentage of loci consistently expressing the opposite strand by tissue type: (BRCA) breast cancer; (LUAD) lung adenocarcinoma; (LUSC) lung squamous carcinoma; (LUCL) lung cancer cell lines; (PRCA) prostate cancer; (PANC) pancreatic cancer; (OV) ovarian cancer; (MENG) meningioma (see Supplemental Fig. S6 for extended version).











