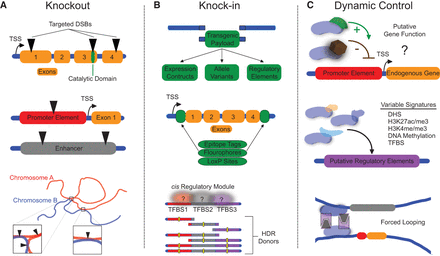
Manipulation of endogenous loci using genome engineering tools. (A) Genetic knockout of coding regions, protein catalytic domains, promoters, enhancers, or genomic contact points is possible using targeted nuclease platforms. (B) Genetic knock-in using appropriate donor repair templates can be applied to deliver various transgenic payloads, including epitope tags, regulatory components, or disrupted motifs such as mutant transcription factor binding sites (TFBSs) within cis regulatory modules, to decipher endogenous regulatory element activity. (C) Dynamic regulation of endogenous loci using nuclease-null genome engineering tools fused to effector domains can be used to interrogate gene function or the activity of putative regulatory elements enriched with varying epigenomic signatures. Additionally, these tools can be used to artificially direct physical interactions between distal endogenous loci. (DSB) double-strand break; (TSS) transcription start site; (HDR) homology-directed repair; (DHS) DNase hypersensitivity site.











