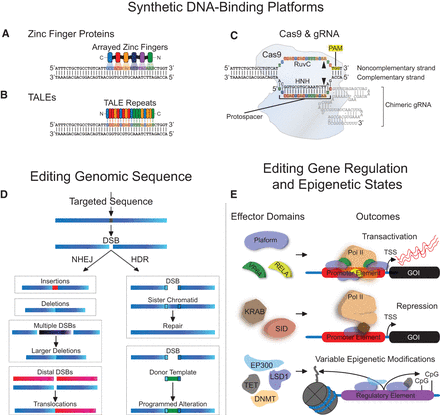
Zinc finger, TALE, and Cas9-gRNA platforms for editing genomic sequence and regulatory states. Individual zinc finger domains (A) and TALE repeats (B) that recognize unique triplets or single base pairs, respectively, can be arrayed in engineered proteins to target specific genomic sequences. (C) Cas9 in complex with a chimeric guide RNA (gRNA) can recognize a specific genomic address through complementarity between the protospacer segment of the gRNA and target DNA. The formation of this complex is dependent upon the presence of a protospacer adjacent motif (PAM). The RuvC and HNH nuclease domains of Cas9 cleave genomic DNA that matches the protospacer (i.e., the noncomplementary strand) and genomic DNA with complementarity to the protospacer (i.e., the complementary strand), respectively (indicated by black triangles). (D) Zinc fingers and TALEs fused to nuclease domains or Cas9 in complex with a gRNA can cleave targeted sequences to generate double-strand breaks (DSBs). DSB resolution through nonhomologous end joining (NHEJ) or homology-directed repair (HDR) can lead to various alterations in genomic sequence. (E) Zinc finger, TALE, or deactivated nuclease-null Cas9 (dCas9) platforms can also be fused to diverse effector domains to modify endogenous gene regulation and epigenetic states: (TSS) transcription start site; (GOI) gene of interest.











