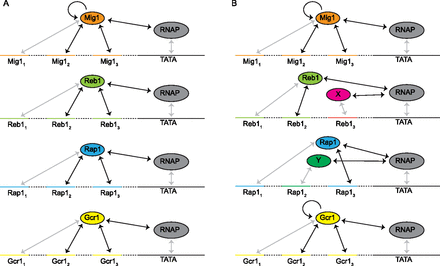
The thermodynamic model consists of a set of interactions that govern TF-DNA and TF-TF binding. Each arrow represents an interaction included in the model in the form of a parameter proportional to the ΔG. Black arrows represent the free parameters. (A) The set of interactions allowed in the simplest model: Each TF is allowed to bind to its cognate TFBSs and to interact with polymerase. Mig1 is allowed to interact with itself when two or more Mig1 sites are present in the same promoter. (B) The set of interactions applied to the model with the highest score: A protein X other than Reb1 is allowed to bind at site Reb13, and a protein Y other than Rap1 is allowed to bind at site Rap12. Both Mig1 and Gcr1 are allowed to interact with themselves when two or more of their sites are present in the same promoter.











