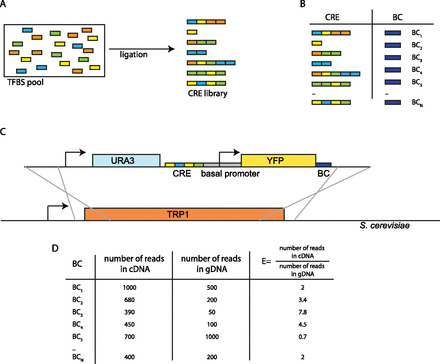
Schematic of the CRE-seq method adapted for this study. (A) Double-stranded oligonucleotides encoding TFBS are mixed in a pool and ligated randomly to create a CRE library. (B) After cloning CRE and barcode (BC) sequences into a reporter plasmid, the concordance between CREs and BCs is determined with a paired-end next-generation sequencing run. Each BC identifies a single CRE. (C) The cassette containing the library of CREs upstream of a basal promoter driving YFP and BC is integrated into the S. cerevisiae genome at the TRP1 locus by selecting for URA+ cells. (D) Cells are grown in liquid culture, and gDNA and RNA are harvested. The fraction of reads in the cDNA pool divided by the fraction of reads in the gDNA pool for each BC is a quantitative measurement of the expression driven by the corresponding CRE.











