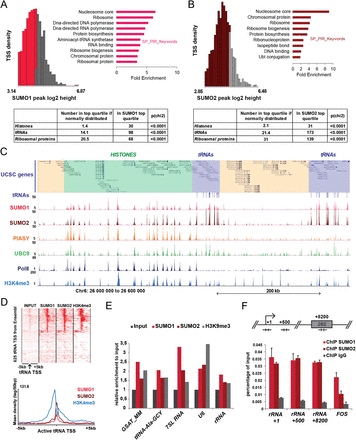
SUMO is highly enriched at histone, ribosomal protein, and tRNA genes. (A,B) Distribution of (A) SUMO1- and (B) SUMO2-marked TSS by peak height and TSS density showing the top quartile corresponding to the highest SUMO peaks in gray. Functional annotation of genes associated to the top quartile TSS using DAVID. The top overrepresented categories are shown. Table below summarizes the statistical enrichment of SUMO-bound TSS for histone, tRNA, and ribosomal protein annotations. (C) A Genome Browser view of ChIP-seq data across the histone and tRNA gene cluster in chromosome 6p. (D) Comparison of SUMO1, SUMO2, and H3K4me3 tag density at tRNA loci (upper) and their merged profiles restricted to the H3K4me3-positive subcluster with respect to distance from TSS (lower). (E) Relative enrichment of SUMO1, SUMO2, and H3K9me3 reads at the major satellite repeat (GSAT_MM), a tRNA, RN7SL1 (7SL), RNU6 (U6), and rRNA loci in the ChIP-seq compared to input. (F) ChIP-qPCR for SUMO1 and SUMO2 on three regions of the rRNA gene amplified with specific primers (Guetg et al. 2012). FOS promoter is used here as a positive control.











