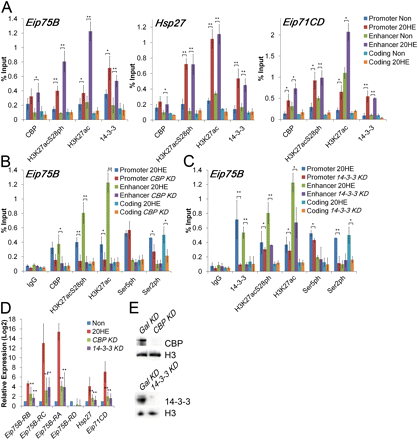
CBP acetylation and 14-3-3 recruitment at promoters and enhancers is necessary for transcription activation of ecdysone-responsive genes. (A) ChIP analysis of Kc167 cells treated for 3 h with ethanol (Non) or 20-HE (20HE) was performed using the antibodies indicated along the x-axis. (Ser5ph) RNAPII phosphorylated on serine 5 of CTD, (Ser2ph) RNAPII phosphorylated on serine 2 of CTD. DNA was quantified by real-time PCR using primers designed to amplify the promoter, enhancer, and coding regions of each of the three ecdysone-induced genes. (B) ChIP analysis of Kc167 cells nontreated and treated with CBP dsRNA compared to Gal dsRNA. The cells were also incubated with 20-HE for 3 h, and chromatin was immunoprecipitated using the antibodies indicated along the x-axis. DNA was quantified by real-time PCR, and the result is reported as a percent of the input at the Eip75B gene. (C) Similar to panel B, but cells were treated with dsRNA to 14-3-3. (D) RNA expression analysis in cells treated with dsRNAs corresponding to the CBP, 14-3-3, or Gal genes. Cells were treated with ethanol (Non) or 20-HE (20HE) for 3 h. RNA levels were determined by qPCR using primers specific to each of the four Eip75B transcripts. All samples were normalized to the mitochondrial gene myt:Col and the Non sample was set to 1 for comparison. (E) Western analysis of lysates from cells treated with CBP (top) or 14-3-3 (bottom) dsRNAs. Levels of either protein are undetectable. Error bars represent the standard deviation of the mean of 3 biological replicates. (*) P < 0.05, (**) P < 0.01.











