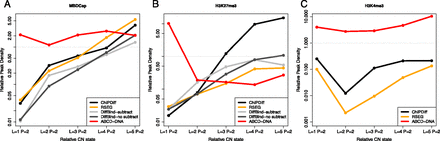
Association between differential peak detection and CNV across LNCaP/PrEC qDNA-seq data sets using various algorithms. The relative rate of peak detection (RRPD), defined as the ratio of the number of regions detected in LNCaP (L) cells to the number of regions detected in PrEC (P), within each CNV stratum is shown for ChIPDiff, RSEG, DiffBind (with and without input subtraction), and ABCD-DNA. DiffBind is based on MACS-detected regions. (A) MBDCap-seq; (B) H3K27me3-seq; and (C) H3K4me3-seq. Due to lack of replication, DiffBind was not run on H3K4me3-seq.











