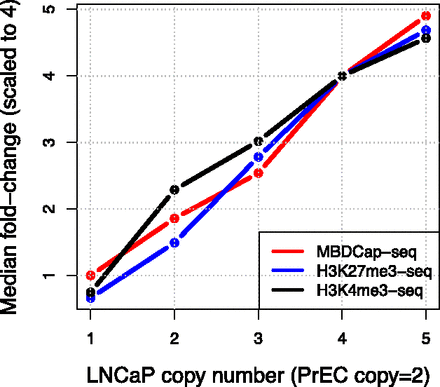
Linearity between CNV and qDNA-seq. Relative read densities scale linearly with CNV for multiple LNCaP/PrEC qDNA-seq (MBDCap, H3K27me3, H3K4me3) data sets. Scaling factors were calculated separately as the median of log-fold changes (median of M-values) for each CNV stratum and each data set (See Supplemental Fig. 3); these medians were exponentiated and scaled according to the most prominent CNV state (L = 4 P = 2). Note that these scaling factors are not actually used in the ABCD-DNA method; they are shown here only to illustrate the relationship between qDNA-seq and CNV.











