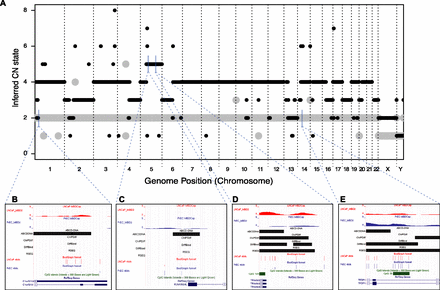
CNV causes false positives and false negatives to various algorithms; ABCD-DNA can recover them. (A) The landscape of CNV between LNCaP (black) and PrEC (gray) cells inferred by PICNIC algorithm (using Affymetrix SNP 6.0 data; see Methods). Using Illumina 450k array data to gauge true differential methylation (see tracks “LNCaP 450k” and “PrEC 450k”), four CNV-induced false-positive (FP) or false-negative (FN) regions in MBDCap-seq data (see tracks “LNCaP_MBD2” and “PrEC_MBD2”) using existing algorithms are shown. Detected differential regions for four methods (ChIPDiff, DiffBind, RSEG, our new approach ABCD-DNA) are shown in black. (B) FN for all algorithms except ABCD-DNA; the change in depth-normalized read density is not particularly strong, but combined with the knowledge that this is a “low” copy region (LNCaP = 2), ABCD-DNA expects fewer reads. Hence, the effective difference is made larger and therefore deemed differential by ABCD-DNA. Similarly, C is amplified in cancer beyond “neutral” (LNCaP = 5), thus ABCD-DNA expects higher read density (if methylated) and correctly increases the effective change. D is similarly amplified, which causes existing algorithms to overstate the differential methylation (i.e., a FP); note the upstream differentially methylated region that all algorithms detect, whereas only ABCD-DNA correctly attributes the downstream change in read density to CNV. (E) Lower copy in LNCaP cells, resulting in lower read depth and FPs for all methods except ABCD-DNA.











