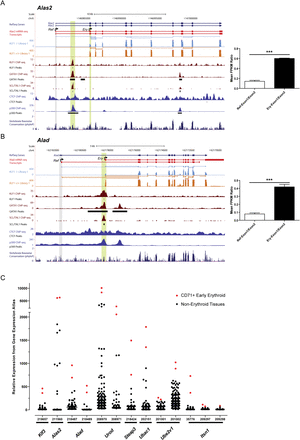
Identification of a novel set of KLF1-GATA1-TAL1–dependent erythroid promoters. (A) An image of murine Alas2 from the UCSC Genome Browser with tracks depicting a novel mechanism of erythroid gene regulation whereby transcription begins at RefSeq exon 2 (Ery TSS). The RefSeq gene annotations (dark blue) are shown together with mRNA-seq transcripts defined using Cuffdiff (red). Representative tracks for a single Klf1−/− and Klf1+/+ mRNA-seq sample are shown in light blue and orange, respectively, with both mapped read density (wiggle) and splice junction information (all tracks are shown in Supplemental Fig. S5). ChIP-seq tracks for KLF1, GATA1, and TAL1 are shown in maroon with peak calls underneath. CTCF and EP300 ChIP-seq from the ENCODE consortium are shown in blue as well as a vertebrate conservation track (phyloP). Gray and green vertical lines are included to point out RefSeq TSSs (Ref) and novel erythroid TSSs (Ery), respectively. Regions of multitranscription factor occupancy are also highlighted by a vertical green line. (Bar chart) Quantification of relative promoter usage describing the ratio of RefSeq exon 1 counts or erythroid exon 1 counts relative to exon 2 for six biological replicates. Data shown are the mean + SEM. (***) P < 0.001 by Student's t-test. (B) An image of murine Alad from the UCSC Genome Browser depicting a novel mechanism of erythroid gene regulation whereby transcription begins at a novel first exon residing within RefSeq intron 1 (Ery TSS). All tracks are shown as above in A. The bar chart also shows quantification of relative promoter usage for either RefSeq exon 1 or the novel erythroid exon 1 compared to exon 2 for six biological replicates. Data shown are the mean ± SEM. (***) P < 0.001 by Student's t-test. (C) Relative gene expression obtained from Gene Expression Atlas for each of the genes described to use novel erythroid promoters (Table 1). Red dots are used to show the expression in the two CD71+ early erythroid samples, while black dots show the expression in all other tissues of the atlas. The HG-U133A probe numbers are shown for each gene since some genes are represented by multiple probes. Klf3 serves as a positive control since it has previously been shown to be ubiquitously expressed with greatest expression in erythroid cells (Funnell et al. 2007).











