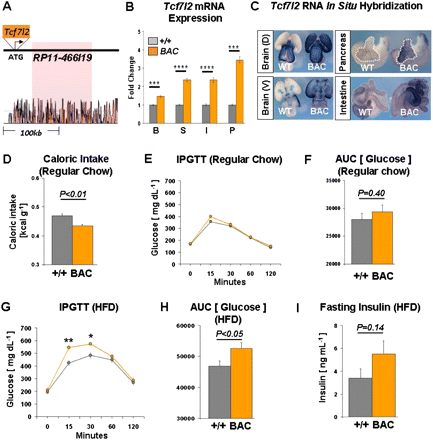
Characterization of Tcf7l2 BAC transgenic mice. (A) Schematic of human BAC RP11-466I19 recombined with a full-length mouse Tcf7l2 cDNA. (Below) Noncoding sequence conservation between human and mouse is marked by red peaks (VISTA genome browser). The T2D-associated interval is highlighted in red. (B) Tcf7l2 mRNA expression from wild-type (gray, n = 4) and Tcf7l2 BAC transgenic (orange, n = 4) mice in brain (B), stomach (S), intestine (I), and pancreas (P). (C) RNA in situ hybridizations of Tcf7l2 BAC transgenic (BAC) and wild-type (WT) tissues at embryonic day 15.5 (E15.5) show overexpression of Tcf7l2 in the brain, pancreas, and intestine in Tcf7l2 BAC transgenic mice. (D) Caloric intake in wild-type (+/+, n = 7) and Tcf7l2 BAC transgenic (BAC, n = 5) mice. (E) Intraperitoneal glucose tolerance test (IPGTT) curves of wild-type (gray, n = 13) and Tcf7l2 BAC transgenic (orange, n = 9) mice on a regular chow diet. (F) Area under the curve (AUC) of the IPGTT plot from E. (G) IPGTT curves of wild-type (gray, n = 8) and Tcf7l2 BAC transgenic (orange, n = 6) mice after 10 wk on a high-fat diet (HFD). (H) Area under the curve (AUC) of the IPGTT plot from G. (I) Fasting plasma insulin levels in wild-type (gray, n = 10) and Tcf7l2 BAC transgenic (orange, n = 8) mice after 10–12 wk on a HFD. (*) P < 0.05; (**) P < 0.01; (***) P < 0.001; (****) P < 0.0001.











