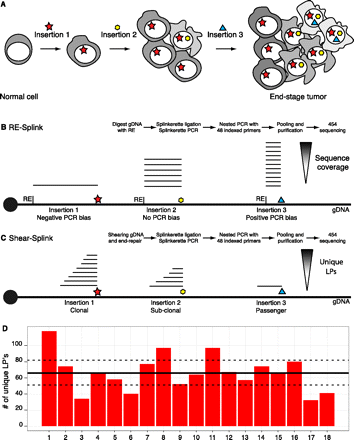
Schematic overview showing the clonality of insertional mutations in tumorigenesis screens, and methods to identify the insertion sites. (A) Clonal expansion of a cell containing an insertion giving a certain growth advantage, which initiates tumorigenesis. In time, additional insertions occur, resulting in a heterogeneous tumor containing a complex collection of insertional mutations. (B) Overview of restriction enzyme based LM-PCR (RE-splink). Amplification of insertion sites using restriction enzymes results in amplicons with a fixed size introducing amplification and sequencing biases, thereby hampering a quantitative identification of insertional mutations within a tumor. (C) Overview of shearing-based LM-PCR (shear-splink), which reduces amplification and sequencing biases and allows the identification of unique ligation points of the splinkerette adapter, each representing a cell within the tumor. (D) Numbers of unique LPs identified for 18 piggyBac insertions in a clonal cell line. The average and 95% confidence interval are indicated by a solid line and dashed line, respectively.











