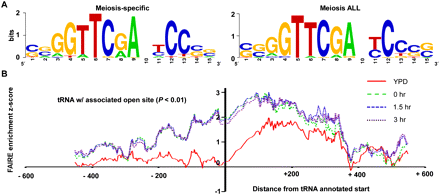
A 15-bp motif is over-represented in meiosis-specific open chromatin surrounding tRNA genes. (A) The 15-bp motif identified from FAIRE-enriched sites isolated from any meiotic sample, but not YPD, and FAIRE-enriched sites isolated from all meiotic samples, but not YPD, are represented (Crooks et al. 2004). (B) Open chromatin increases during meiosis at sites associated with tRNA genes. The position relative to tRNA annotated start sites (base pair) is plotted on the x-axis, and FAIRE enrichment is plotted on the y-axis. Cytoplasmic tRNA genes associated with a site of open chromatin were aligned at their annotated transcription start. Shown is a moving average of the average FAIRE-enrichment z score (window size = 15 probes). FAIRE data from YPD (red), 0-h (green dashes), 1.5-h (blue dashes), and 3-h (purple dots) samples are plotted. Regions corresponding to the functional tRNA have a median length of 72 bp (minimum of 70 bp and a maximum of 132 bp). The graph includes the 98 tRNA-associated sequences that overlap a site of open chromatin (P < 0.01).











