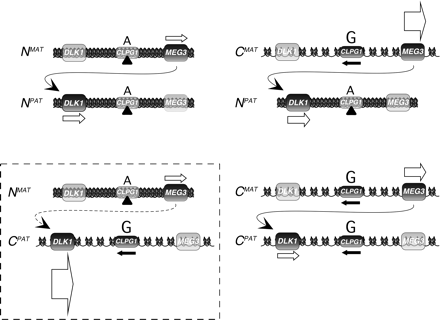
Mechanistic model for the callipyge mutation. Schematic representation of the four genotypes at the callipyge locus in adult sheep, the patterns of gene expression for DLK1, MEG3, and CLPG1, and a model for chromatin structure of each allele based on our current and prior findings (Murphy et al. 2005). The relative position of the mutation within the CLPG1 locus is shown above each allele (A, normal; G, mutated). Arrows beneath transcribed DLK1 and MEG3 are proportional to transcription levels (Murphy et al. 2005). In adult sheep, the ratio of DLK1:MEG3 transcripts is highly correlated with phenotype in a temporal and tissue-specific manner, suggesting that increased MEG3 transcription is necessary for the postnatal down-regulation of DLK1. In normal sheep, attenuation of DLK1 transcription is accompanied by a doubling of cytosine methylation in the region of the callipyge mutation. Reciprocal heterozygotes and CMATCPAT exhibit deficient cytosine methylation, and this may reflect a more open chromatin structure and permissiveness for transcription. This is supported by the observed monoallelic expression of CLPG1 only from alleles carrying the mutation (black arrows). The mutation may therefore alter the ability of regional chromatin to condense perinatally via inhibition of chromatin remodeling complex assembly (black triangles), inhibition of DNA methylation, and/or altered histone modifications.











