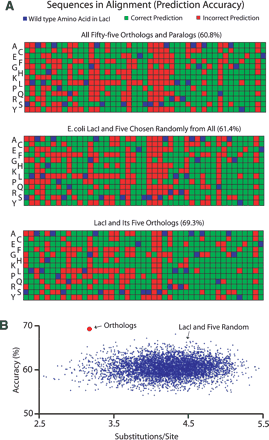
Effect of paralogous sequences on prediction accuracy. (A) Differential accuracy in MAPP classification of LacI variants implicated in ligand binding when different types of homologs are represented in the alignment. Three types of alignment are compared. Variants classified at each position are arranged from top to bottom alphabetically by one-letter abbreviation. Positions are shown left to right in increasing order (24). Green and red identify correct and incorrect predictions, respectively, of whether a variant is functional versus deleterious; the wild-type amino acid is blue. (B) Classification of 5000 alignments, each containing LacI and five sequences randomly chosen from the original alignment. Accuracy is plotted against total evolutionary divergence as measured in substitutions per site for random alignments (blue) and the single alignment of six orthologs (red).











