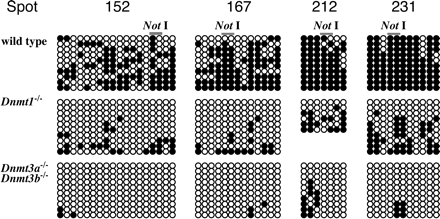
DNA methylation status at CpG sites around the NotI sites. Methylation status around ∼600-bp genomic region, including the NotI site, was assessed by sodium bisulfite sequencing method. Open circles and closed circles represent unmethylated and methylated cytosine residues, respectively. In wild-type ES cells, all genomic regions are hypermethylated, including the NotI sites, indicating that DNA methylation is maintained at these regions in the wild-type ES cells. In contrast, CpG sites in the same regions are demethylated in Dnmt1-/- and Dnmt3a-/-Dnmt3b-/- ES cells. Although the demethylation is moderate in Dnmt1-/- ES cells, almost all CpG sites are totally demethylated in Dnmt3a-/-Dnmt3b-/- ES cells, indicating that RLGS analysis reflects DNA methylation status not only at the NotI site but surrounding CpG dinucleotides of the NotI site. These data suggest that Dnmt3a/3b are more significant for DNA methylation in CpG islands than Dnmt1.











