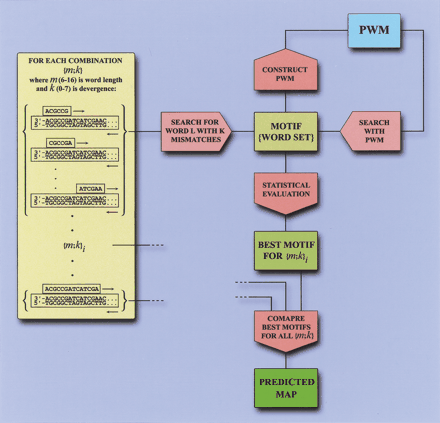
Scanseq algorithm. Initial search is performed with words of length m with 0-k mismatches. For each word found in the sequence, the corresponding motif (word set), is refined by positional weight matrix (PWM), and is statistically evaluated through Z score. In the final stage, Z scores for motifs within a range ofm and k are compared and a predicted map is generated. Note that the PWM in the Scanseq algorithm is not the same as in the strategy of BSTF map construction, and it does not include any a priori information about binding motifs.











